Last modified 2026-04-10 |
|
Use the Read Files SDK Method (Tutorial)
At a Glance
This document explains how to use read_files() to download files to your HISE IDE. If you have questions, contact Support.
Method signature
read_files()
hp.read_files(
file_list: list = None,
query_id: list = None,
query_dict: dict = None,
to_df: bool = True,
is_public: bool = False,
)readFiles()
readFiles(
fileIds = list | NULL,
queryId = character(1) | NULL,
query = list | NULL
isPublic = bool | FALSE,
)Parameters
The parameters for this method are listed in the following table. In each key:value pair, the value must be of type list.
Python Parameters | ||||
Parameter |
| Data type |
| Description |
|
| list |
| List of UUIDS to retrieve |
|
| string |
| Value of the queryID from an advanced search |
|
| dict |
| Dictionary that allows users to submit a query |
|
| bool |
| Boolean determining whether the result is returned as a DataFrame |
| bool | Boolean determining whether to query public files or not | ||
R Parameters |
| ||
Parameter |
| Description |
|
|
| List of UUIDS to retrieve |
|
|
| UUID from an advanced search |
|
|
| List of query params to search for. The format is similar to that passed to getFileDescriptors, but the fields correspond to fields in the Subject materialized view. NOTE: fileType with a valid entry must be present |
|
| Boolean determining whether to query public files or not. | ||
Description
This function fetches HISE files and returns one or more objects when you pass in the following:
- A list of file IDs (Python:
file_list| R:fileIds) - A saved search ID (
query_id| R:queryId) - A custom search query (
query_dict| R:query)
The object returned is either a dictionary (Python) or a list of data.frames (R). The dictionary or list contains keys [descriptors, labResults, specimens, values].
Instructions
The following instructions are written for Python. To adapt them for R, use the R function signature and parameters listed above.
 Import libraries
Import libraries
To get started, set up your environment to interact with HISE programmatically and access all available SDK functions. For details, see Use Hise SDK Methods and Get Help in the IDE.
1. Navigate to HISE, and use your organizational email address to sign in.
2. Open an IDE. For instructions, see Create Your First HISE IDE (Tutorial).
3. For programmatic access to HISE functions and efficient handling of tabular data, import the Python SDK and the pandas library.
# Import hisepy and pandas
import hisepy as hp
import pandas as pd
 Define file IDs
Define file IDs
In this step, we define the file IDs for this notebook. For details, see Use Advanced Search (Tutorial).
1. Retrieve your own set of file IDs, and then define them as shown below. (The example below uses placeholder UUIDs—replace them with your own.)
# Define the file IDs used in this analysis
FILEIDS = ['4551e620-48db-4328-a2b0-122730cd128d', '6417a4c5-098b-4d70-8c24-951e1c1c44ce']
 Return dictionary output and apply tabular format
Return dictionary output and apply tabular format
| To see what's in a given dictionary key, use the following format:
|
When you call read_files() with the to_df=True parameter, a dictionary is returned in which each key contains a pandas DataFrame. The to_df=True parameter arranges the data into a tabular format for easier analysis.
1. Pass your list of file IDs to read_files().
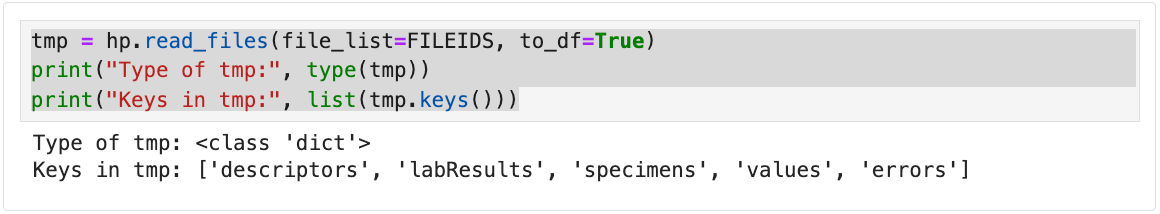
# Return dictionary output and print keys from read_files
tmp = hp.read_files(file_list=FILEIDS, to_df=True)
# Shows the class of the returned object
print("Type of tmp:", type(tmp))
# Prints all keys (file IDs or names) in the dictionary
print("Keys in tmp:", list(tmp.keys()))
The following output is returned.
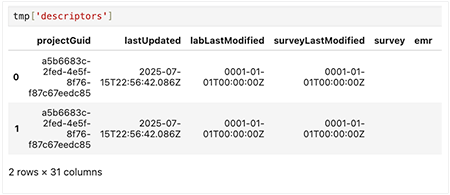
 Preview the data
Preview the data
Each key in the tmp dictionary represents a different dataset returned by hp.read_files(). The accompanying table summarizes the content of each key.
| Key | Description |
|---|---|
descriptors | Project, sample, or subject metadata |
labResults | Test results and IDs |
specimens | Status and info on biological specimens |
values | Raw data metrics |
errors | File retrieval errors, if any |
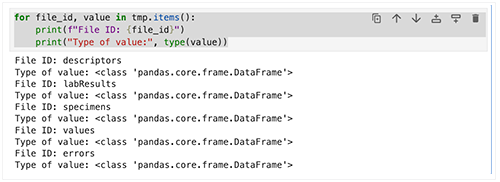
1. For each key, use a loop to print the file ID, the value type, and a preview of the data.
for file_id, value in tmp.items():
print(f"File ID: {file_id}")
print("Type of value:", type(value))
# If it's a DataFrame, show the first few rowstry:
print(value.head())
except AttributeError:
print(value) # For non-DataFrame types
print("-" * 40)
2. To see all column heads for a given data set, use the following line.
print(tmp['descriptors'].columns)
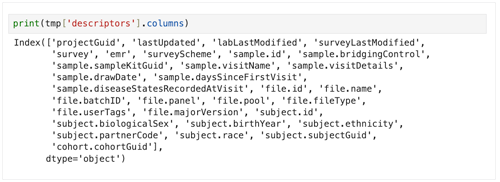
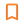
3. To get a summary of the DataFrame, use the following line.
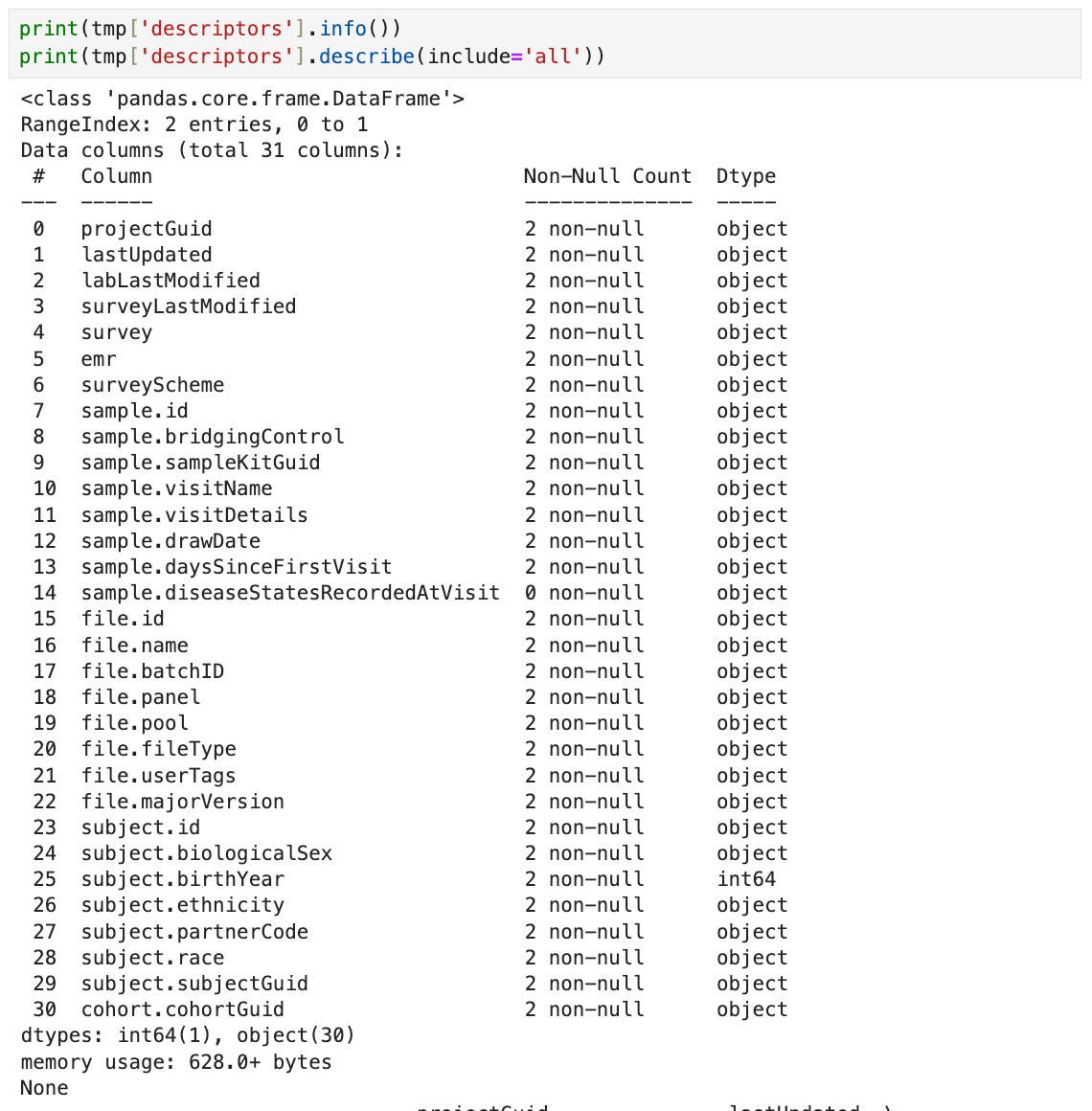
print(tmp['descriptors'].info())
print(tmp['descriptors'].describe(include='all'))
 Related Resources
Related Resources
Create Your First HISE IDE (Tutorial)
Prepare your visualization in an IDE
Navigate to HISE, and use your organizational email address to sign in.
Open an IDE. For instructions, see Create Your First HISE IDE (Tutorial) .
To import
hisepy, enter the following code into a new cell in your IDE.
# Import the Python SDK to enable programmatic access to HISE functions
import hisepy as hp4. Load your data and generate a visualization, according to the needs of your study.
A. For simple visualizations, create a Plotly figure.
i. To make sure your code runs end to end in the IDE, run the following code in a new IDE cell:
fig.show()
ii. To save the visualization, store it in a Plotly figure object to pass in later, as shown in the following block:
fig = px.scatter(
x=[1, 2, 3, 4],
y=[10, 15, 13, 20],
title="Plotly example"
)
# Create scatter plot
# Define values on x axis
# Define values on y axis
# Set chart titleB. For complex visualizations, build an app (for example, a DeckGL app) in your IDE workspace (for example, /home/workspace).
Create and save a hero image
Each saved visualization in HISE includes a static image visible to users who are browsing among the saved visualizations.
From your IDE, export or otherwise save a PNG that represents your visualization (for example, a Plotly figure export or a screenshot of your app’s main view).
Save the image somewhere in your IDE workspace (for example,
/home/workspace), so the SDK can access it.
Path 1: Save a Plotly visualization As a starting point, you can use the sample Plotly notebooks in the IDE’s examples folder.
Import the HISE SDK module.
import hisepy as hpImport Plotly Express.
import plotly.express as pxConfirm and note the file path to this PNG. You’ll need this path later, when you make the SDK call to save the visualization.
fig.write_image("/home/workspace/example-plot.png")Call the SDK. Call
save_visualization()with its required and optional parameters. The SDK saves your Plotly figure into the selected study as a saved visualization. HISE links that saved visualization to the IDE run that produced it, including input data, code, and environment.hp.save_visualization(figure=fig, studyspaceid="12345678-1234-1234-1234-123456789abc",title="Example Plotly visualization",png_image="/home/workspace/example-plot.png")# SDK call# Plotly figure object# Study space ID# Descriptive title# Path of hero image
Path 2: Save a visualization app Use this path when you’ve built an app (for example, using DeckGL or Dash) and want HISE to build and deploy it using a visualization build template.
Note your file path. Place your app files and directories in a location HISE can package. You’ll pass in these file and directory paths to
save_visualization_app()./home/workspace/my-appInclude your app’s main entrypoint and any framework-specific files (for example, DeckGL,
package.json, server entry files).Any necessary configuration files or scripts for your app (for example,
requirements.txt, config scripts).Do not include large input data files. Instead, upload those separately to your study space for your app to use when you run it.
Capture your input file IDs. Before you call
save_visualization_app(), save your input data files in HISE, since the app must read from those files.If your app uses files already present in your study space, capture their file IDs from HISE and have them ready to pass in.
If your app uses files generated in the IDE that aren’t yet in HISE, upload them using the
upload_files()(Python) |uploadFiles()(R) SDK method, and wait until those files become available.Collect the file IDs you want to include in your app, and keep them in a list (for example
data_source_fileids()). You pass in these IDs to thedata_source_fileidsparameter.Select the data mount path. Choose a path (for example
/dataor/input) as the directory in which your input data is stored, so your app can read it at runtime. You pass in this path as thedata_mount_pathargument. If you forget the leading/, HISE adds it for you.Import the HISE SDK module.
import hisepy as hpCall the SDK. Call
save_visualization_app()with the required parameters:
hp.save_visualization_app( application_files=["/home/workspace/my-app/app.py", "/home/workspace/my-app/requirements.txt"], application_dirs=["/home/workspace/my-app/configs"], study_space_id="12345678-1234-1234-1234-123456789abc", title="Example visualization app", png_image="/home/workspace/example-app.png", data_mount_path="/data", data_source_file_ids=[ "9f6d7ab5-1c7b-4709-9455-3d8ffffbb6c8", "0fb06e51-74c4-46be-b92d-5e045232b2d9" ] ) |
List of app file paths App directories Study space ID Title (10+ characters) Image path (PNG) Where app reads data Input data IDs for files in HISE */ |
|---|
(Optional) If you already know which visualization build template you want to use (for example a specific DeckGL template version), you can also pass in the following additional parameters:
i. build_template_name
ii. build_template_major_version
iii. build_template_minor_version
iv. build_template_parameters /* A map of framework-specific arguments, such as app entrypoint paths, which must match the build variables for that visualization build template /
v. infer_build_template_arguments /* Parameter that tells HISE whether to infer framework-specific arguments from your files and directories if you don’t supply them directly */
The save_visualization_app() SDK then packages your app files, applies the selected visualization build template, and starts a build workflow to deploy your app. It also creates a saved visualization in your study that points to the deployed app.
Call the SDK
Review the saved visualization in HISE
Find the saved visualization, and share it with your team.
From the top navigation menu, click COLLABORATION SPACE.

Click the card that represents your study.
In the side navigation menu, click Visualizations.
Find the card that represents your saved visualization (the one with your hero image on it), and click to open it.
Review your saved Plotly figure or app-based visualization in Draft mode.
If you like, confirm the output with your collaborators. For instructions on adding collaborators to your study, see Work with Studies (Tutorial) . When you're satisfied with the saved visualization, mark it as Final.