Last modified 2026-02-06 |
Use the Read Subjects SDK Method (Tutorial)–DRAFT
 | Abbreviations Key | ||||
| HISE | Human Immune System Explorer | SDK | software development kit | ||
hp | hisepy | ss | study space | ||
| IDE | integrated development environment |
At a Glance
This document explains how to use read_subjects()(Python) | readSubjects(R) to retrieve subject-level metadata you can filter or join with other analysis results.
The read_subjects() signature is shown in the box below, and the method parameters follow. You must specify either the subject ID or the query parameter. If you have questions or need help, contact Support.
Method signature
 read_subjects()
read_subjects()
hp.read_subjects(
subject_ids: str = None,
query_dict: dict = None,
to_df: bool = True,
) readSubjects()
readSubjects()
readSubjects(
subjectIds = NULL,
query = NULL,
toDF = TRUE, ...
)
Parameters
The parameters for this method are listed in the following table. In each key:value pair, the value must be of type list.
Python Parameters | |||
| Parameter | Data type | Required or optional | Description |
subject_ids | str | required | A list of Subject UUIDs or a single query ID |
query_dict | dict | required | A dictionary object containing search parameters using Mongo query language. |
to_df | bool | required | If true, returns a data.frame. Default is True. |
R Parameters | |||
| Parameter | Data type | Required or optional | Description |
subjectIds | list | required | A list of Subject UUIDs or a single query ID |
query | list | required | A list of query params to search on. The format is similar to that passed to getFileDescriptors, but the fields correspond to fields in the Subject materialized view. |
toDF | bool | required | Boolean indicating whether the output should be reformatted to a data.frame. Default is True. |
 Get Help
Get Help
If you get stuck during a read_subjects() | readSubjects call, refer to the steps of this tutorial (examples are in Python unless otherwise specified). To use the baked-in help in your IDE, try one of the following commands. Still not working? To file a ticket, click Collaboration Space. Doing so lets you share your insights with others. The read_subjects() | readSubjects signature is shown in the box below, and the method parameters follow. If you have questions, contact Support.
| Python | R | Output |
help(hp.read_subjects) | help(readSubjects) | Function signature, list of parameters, class, and a brief description of the method in a compact plain-text format |
hp.read_subjects? | ?readSubjects | Method signature, docstring (description), file location, and file type in more readable format |
hp.read_subjects?? | hise::readSubjects | Signature, docstring, file path, a verbose set of metadata, and the source code for the method |
Instructions
 Import libraries
Import libraries
To get started, set up your environment to interact with HISE programmatically and access all available SDK functions. For details, see Use Hise SDK Methods.
1. Navigate to HISE, and use your organizational email address to sign in.
2. Open an IDE. For instructions, see Create Your First HISE IDE (Tutorial).
3. To import hisepy, enter the following code into a new cell in your IDE.
# Import the Python SDK to enable programmatic access to HISE functionsimport hisepy as hp
# For downstream analysis (optional)import pandas as pd
 Retrieve either subject IDs or a query dictionary
Retrieve either subject IDs or a query dictionary
You must provide either a list of subject IDs or a query dictionary, but not both.
Subject IDs
# Pass in subject IDs
subject_ids = [
"3c2514cf-9b7e-4b68-a11b-81cc46aa3985",
"1b8a6f6e-1234-4abc-9def-0123456789ab",
]
# Pass in a query
query = {
"cohortGuid": "FH1",
# Include additional fields as needed
}
 Call
Call read_subjects()
Pass your chosen parameter to hp.read_subjects(), and optionally control whether the result is returned as a DataFrame.
# Example: Retrieve specified subjects by ID
subjects_df = hp.read_subjects(
subject_ids=subject_ids,
query_dict=None,to_df=True,
)
# Example: Search for subjects with a query
subjects_df = hp.read_subjects(
subject_ids=None,
query_dict=query,
to_df=True,
)
If to_df is False, read_subjects() returns the JSON payload instead of a DataFrame.
 Track the status of your call
Track the status of your call
1. To track the status of a read subjects call, check the IDE for an error message or success notification, as shown in the accompanying image.

2. For detailed information, return to HISE:
A. Under RESEARCH, click the card for your IDE. A tag indicates whether the call has succeeded, failed, or is still in progress, and the Upload file logs show the status of all recent calls.
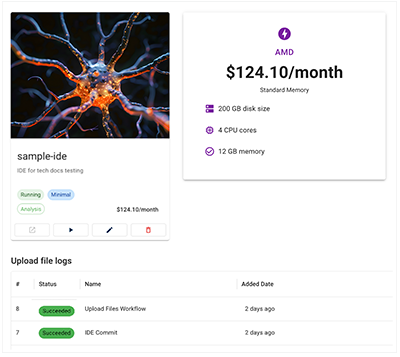
B. If you used Fast Mode to upload files to an IDE, a small blue tag appears next to the IDE in the list. For details, see Use Fast Mode (Tutorial).

C. For granular information about your file uploads, check the IDE Instances page in the COLLABORATION SPACE. It displays an upload files workflow banner, list view, and detail view. For details, see Work with Studies (Tutorial).
 Work with the results
Work with the results
After you retrieve the subject metadata, you can inspect and filter it like any other Pandas DataFrame.
1. To preview the data, use the following command:
# Preview columns and a few rows
subjects_df.head()
# Example: Filter by diagnosis or cohort
fh1_subjects = subjects_df[subjects_df["cohortGuid"] == "FH1"]
 Related Resources
Related Resources